 The numbers of tra genes on circular plasmids of Streptomyces are small (usually one or two) compared with those on circular plasmids of Gram-negative bacteria (often> 10). Genes involved in intermycelial transfer ( tra/ kil genes) and intramycelial spread ( spd genes) during conjugation have been identified in several circular plasmids. Like their circular counterparts, the linear plasmids of Streptomyces also mediate conjugation and promote the transfer of themselves and the donor chromosomes (reviewed by Hopwood and Kieser, 1993). These palindromic sequences may form complex secondary structures that are involved in the patch-type replication of the 3′ overhang ( Huang et al., 1998 Qin and Cohen, 1998). The terminal sequences of many linear Streptomyces plasmids consist of several densely packed palindromes that share extensive homology with those of the linear chromosomes for about 200 bp ( Huang et al., 1998). In at least two linear plasmids (pSCL and pSLA2) investigated, replication has been shown to be initiated internally and proceeds bidirectionally toward the ends, leaving single-stranded overhangs at the 3′ ends ( Chang and Cohen, 1994) to be patched probably by synthesis primed by TP. The size of these linear plasmids varies widely (from about 10–1000 kb), and so does the size of their terminal inverted repeats (TIRs 44 bp to about 80 kb). The soil bacteria Streptomyces are unusual among eubacteria in having terminal protein (TP)-capped linear chromosomes ( Lin et al., 1993) and plasmids ( Sakaguchi, 1990). 
#MACVECTOR PLASMID MAPPING PLUS#These, plus the inferred recombination at the right arm, indicate that SLP2 has evolved through rounds of exchanges involving at least three replicons. The G+C content and GC or AT skew profiles exhibit complex distributions. In addition, two pseudotelomere sequences are present near the left telomere. 
lividans using the chromosomally encoded terminal protein. Mini-linear plasmids containing these telomeres replicated in S. The two telomeres diverge significantly in primary sequence, while preserving similar secondary structures. The terminally located helicase-like gene ttrA was necessary for conjugal transfer. The 43 putative protein coding sequences contained many involved in replication (including two terminal protein homologues), partitioning, conjugal transfer and intramycelial spread. Examination of the flanking target sequences of Tn 4811 suggests a previous recombinational event there. The rightmost 15.4 kb sequence is identical to that of the host chromosome, including the Tn 4811 sequence at the border, which is interrupted by an insertion sequence (IS) element in SLP2. The nucleotide sequence of SLP2 was determined. Terms of Use.SLP2 is a 50 kb linear plasmid in Streptomyces lividans that contains short (44 bp) terminal inverted repeats and covalently bound terminal proteins. So, for example, you could adjust the sliders so that only enzymes with no cuts are shown in the list, then save those to a file, then use that file in a restriction enzyme analysis of a second sequence to identify enzymes that cut that, but that do not cut the first sequence.Ĭopyright © 2022 MacVector, Inc. At any time you can save the current set of enzymes to a separate. The Filter controls are dynamic, so that as you drag the sliders, the list of enzymes updates to reflect only those with the appropriate number of cuts, appropriate site size and end structure. The RE Picker window consists of an upper area with a variety of controls and a lower area with a list of enzymes A similar button controls the visibility of the floating Graphics Palette. The floating RE Picker is quite a large window in order to provide a reasonable visible list of enzymes, so there is a convenient button on the Map tab toolbar to show/hide the window so that it is only visible when you are interacting with it. all the non-cutters) for use in subsequent analyses. This gives you very fine control over the enzymes that are shown and you can save sets of enzymes (e.g. However, all of the sites that meet the cut criteria are displayed in the RE Picker, and there are a number of sliders and controls that let you filter the enzymes displayed based on number of cuts, site size, cut site overhang type and selection status. By default, each time you open a DNA sequence, MacVector scans the sequence for ALL of the restriction sites in the selected restriction enzyme file, but only displays those that are selected in the file. Introduced in MacVector 17, the RE Picker lets you dynamically filter the enzymes displayed in the Map tab of DNA sequence documents. 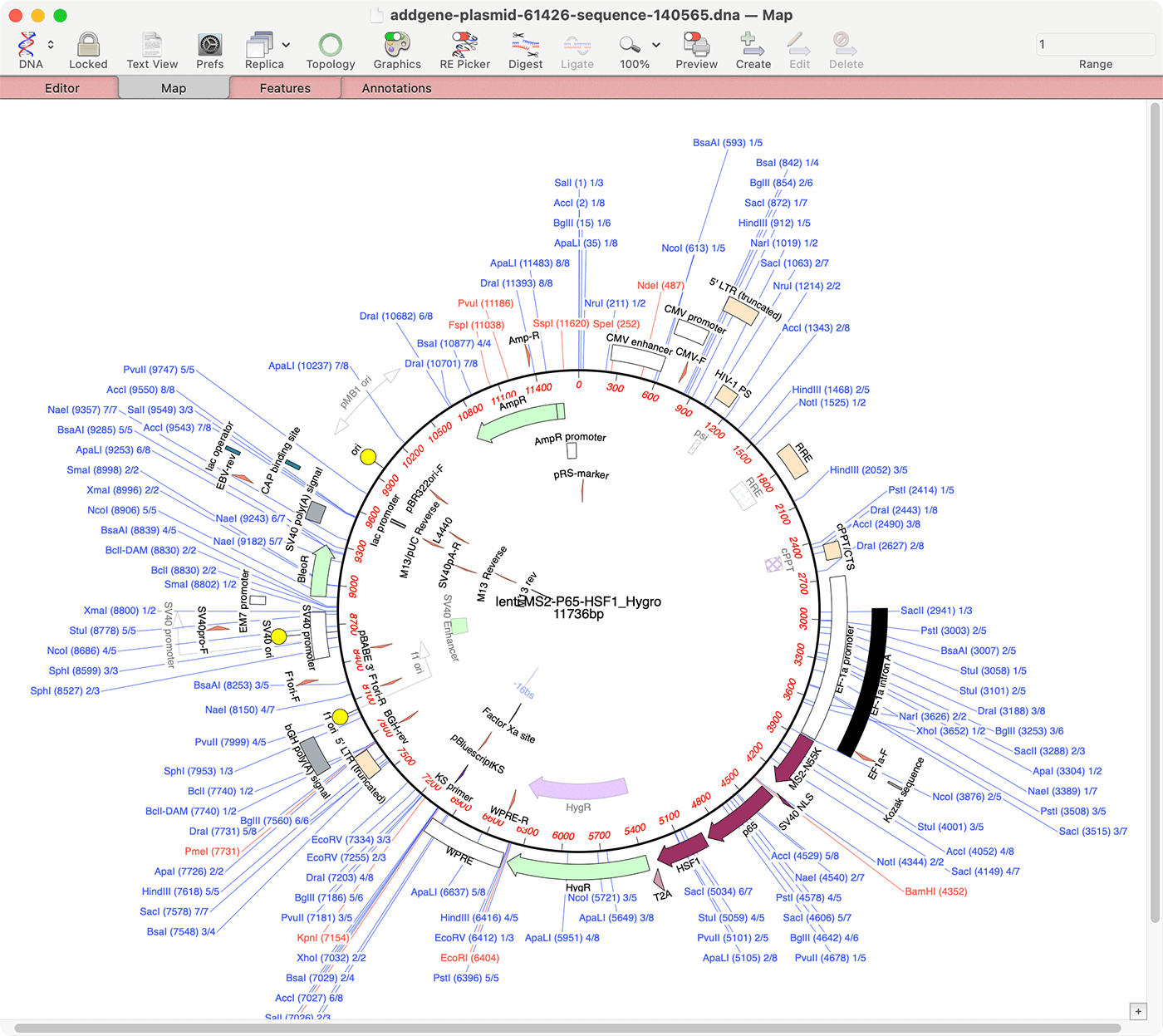
Interactive Restriction Enzyme Picker (RE Picker) Sequence Analysis Tools for Molecular Biologists 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
